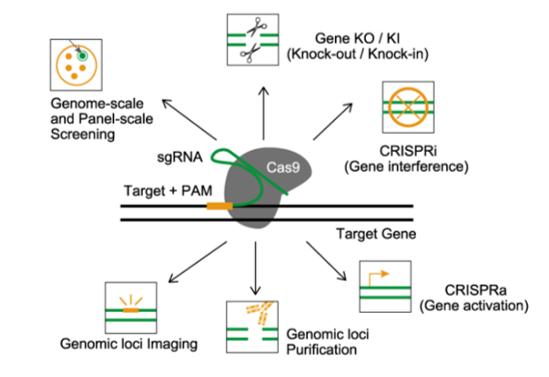
Synbio Technologies has extensive experience with gene/genome editing. We can provide one-stop services ranging from single sgRNA/sgRNA library design/synthesis and lentivirus packaging to bioinformatics analysis and much more. By relying proprietary gene synthesis platform, we have successfully completed efficient and high-throughput gene/genome editing.
Why Us?

Database covers sgRNA design & synthesis for different species, including animals, plants, microorganisms.
Professional staff offer customized design of sgRNA for gene family or genome region, without gene sequence or species restrictions.
We provide comprehensive services, including Ready-to-use sgRNA, plasmid of sgRNA, and sgRNA libraries.

We provide professional single sequence and sgRNA library design of various species for gene knockout, repression and activation.
High throughput and most cost-effective.
DNA synthesis platform, starts at $0.09/bp for synthesis.
shRNA, TALEN, CRISPR-Cas9 vector construction and validation.
High quality standard present reliable gene sequencing and analysis services.
Guaranteed 3 mg protein of 85% purity with our Syno® Gene to Protein Package in E. coli expression system.