Synbio Technologies provides professional sgRNA bioinformatics services for CRISPR-Cas9 system. The services we offer range from sgRNA design, synthesis and gene editing, to gene expression profiling analysis to chromosomal region with DNA copy number statistics based on sequencing results as functional analysis. Combined with professional databases such as PRECOG, TCGA and COSMIC, we can analyze gene expression levels, gene mutation rates and relevance of clinical prognosis. This allows us to provide solutions for potential drug targets and personalized medicine. It could also help scientists find more genes serve as therapeutic targets.
One-Stop Service for sgRNA

sgRNA Bioinformatics
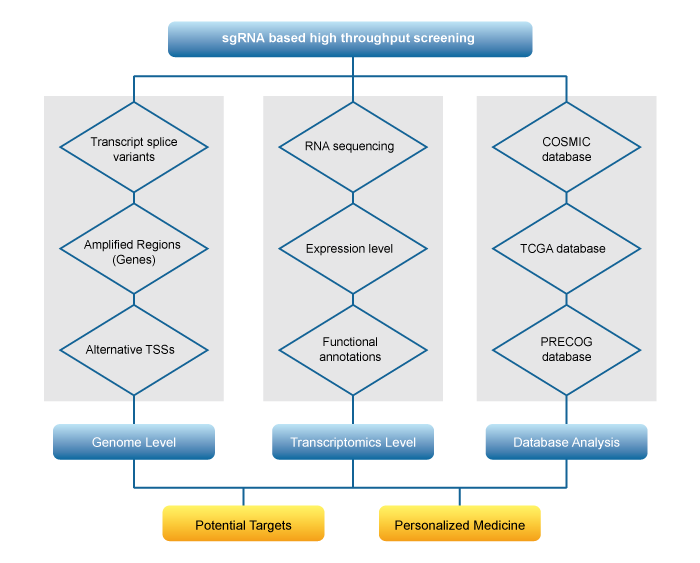
Abbreviation:
Alternative TSS: Alternative Transcription Start Sites
Competitive Advantages
- Experienced in >20 species for CRISPR-Cas9 sgRNA design, which can be customized for specified requirements.
- Professional bioinformatics analysis team with scientific scheme, in-depth data mining to generate a reliable and flexible data report.
Applications
Case Studies
The cells of different samples were infected with viruses carrying CRISPR-Cas9 sgRNA library. The cells were then sequenced with next-generation sequencing technology.
We first evaluated the effect of sgRNA knock out. Secondly, the enriched or depleted sgRNAs were analyzed and compared to screen the genes and targets for cell survival or drug action by functional annotations such as GO and KEGG. Gene Set Enrichment Analysis is the process of finding a class of genes or proteins that are overexpressed in a group of genes or proteins. The common ones are GO functional annotation and KEGG pathway enrichment analysis. Through gene pathway enrichment analysis, we can preliminarily analyze the biological processes or signaling pathways that differentially expressed genes may participate in. Finally, the targets were further analyzed by combining the cancer database and mutation database.
The results are as follows:
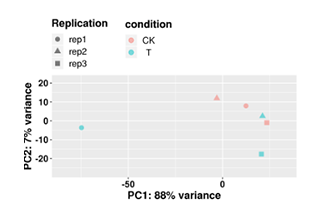
PCA Analysis
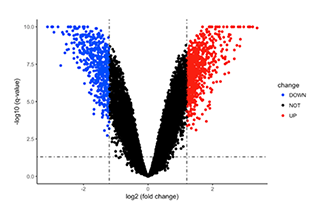
Differentially expressed gene volcano plot
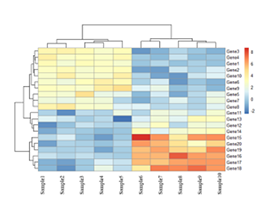

Effect of sgRNA knock-out on gene expression

Differentially expressed gene venn diagram
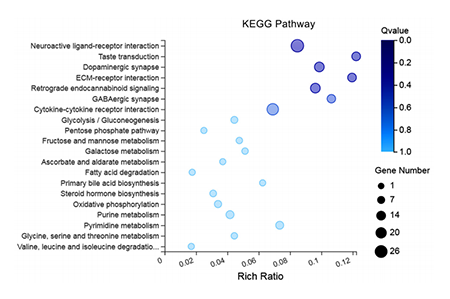
KEGG pathway enrichment plot
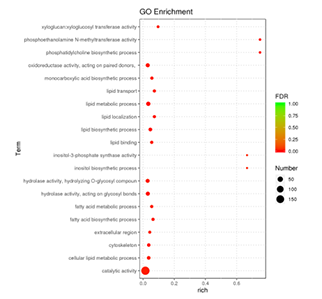
GO enrichment bubble plot
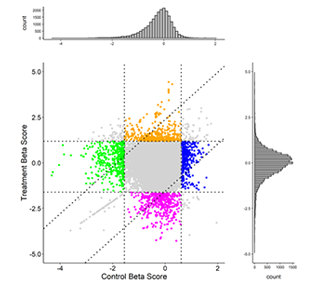
Scatter plot of beta score
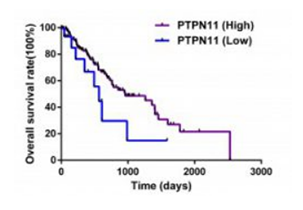
Mining potential drug targets with data from TCGA database
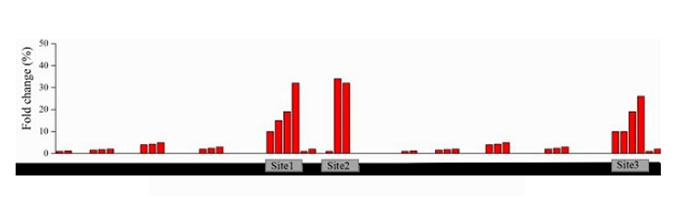
Analysis of sgRNA on the knock out sites of exons
Service Specifications
| Service | Analysis | Analysis Software | Turnaround Time | Deliverables | Price |
|---|---|---|---|---|---|
| sgRNA Bioinformatics Analysis | Genome, Transcriptome, Drug Target Mining, etc. | R software | 7-10 Business Days | Analysis Report | Quote |

